Albert-László Barabási is the Robert Gray Dodge Professor of Network Science at Northeastern University, and he also holds an appointment in the Department of Medicine at Harvard Medical School. We talked about a revolutionary ‘network medicine’ approach that can greatly enhance our ability to understand biological processes and seek cures for disease, and we also discussed why ultra-processed food is really, really bad for you.
You’re a physicist by training, which is pretty cool if you ask me. How did you end up studying biology and specifically the biology of aging?
Almost by accident. I got interested in networks around 1994-95. I was living in Chicago at that time, and I met a medical researcher. We started talking, and he said he was studying cancer. I said, I study networks, and he said, you know, there are lots of networks in biology. So, he asked me to tell him more. This conversation continued, and he actually found us a network, the metabolic network, to work on. We wrote a great paper about it, sent it to Nature, and it was accepted.
Then, he brought us another network that he called the protein interaction network. We analyzed it, wrote another paper, sent it to Nature, and it was accepted again. After that second paper, I thought maybe I should actually learn a bit about cell biology, and that’s how my journey began. I got a big book, sort of the Bible of cell biology, started to read it, and I said, geez, if only I would have known all these details about how the metabolism network and the protein interaction network work, I would have probably never dared to write those papers. Ignorance is bliss, right?
Soon after, I came to Dana-Farber Cancer Institute at Harvard Medical School. I spent a year there on sabbatical, one thing led to another, and now I’m in the medicine department, among others, at Harvard.
What was your opinion of biology as science when you encountered it?
I first encountered biology in high school, and for me it felt like a phone book where you had to learn lots of terms that you didn’t really understand and you also didn’t understand how they all fit together. That’s not a criticism of biology, more like of how biology was taught in Transylvania, Romania, where I’m from, back when I was a student.
As a result, I had absolutely zero interest in biology, and I only reconnected with it in a meaningful way when I was at Dana-Farber. That was really exciting time because, by then, biology became digital. This was the time, around 2005-2006 and just a few years after the Human Genome Project, when everybody around me was discovering yet another gene connected to yet another disease. It became a digital story: there were units that were connected in a discreet way.
For me, it was a game changer in how to approach it. Before that, with my colleague Zoltan Oltvai, we’d be focusing on model organisms, but when I went to Dana-Farber, I realized there’s so much more interesting data on humans, so I switched my lab entirely away from model organisms and straight into human biology. It’s a journey that has never ended, as we’re continuing to focus mainly on humans.
Human biology is, of course, a giant field, and many people are contributing to it greatly. What we brought to this discussion, however, is the network perspective, and what had started at Dana-Farber eventually converged into the network medicine division at Harvard, where today, more than 200 medical doctors and researchers are focusing on the network paradigm.
I’m not sure it’s even doable in the format of an interview, but we have to try. Could you explain what network science is and why do you think it is a proper way to study living things?
Let’s start with network science in general. By the way, my lab is bigger than biology; it focuses on networks in general. Network science is important because most complex systems around us work as networks, and we cannot understand them without mapping the networks around.
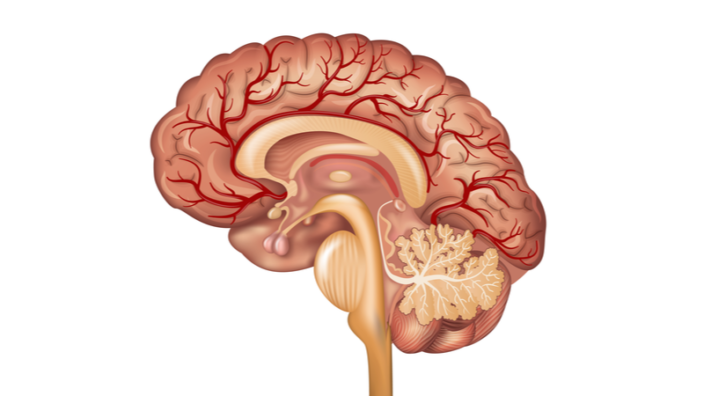
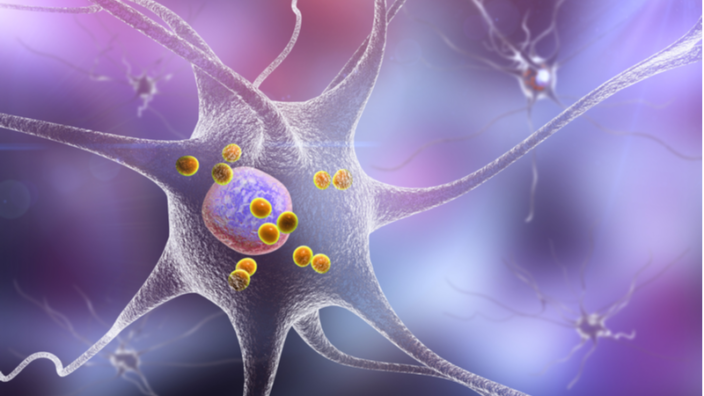
The most obvious example is the cell, which is a network of chemicals, proteins, and metabolites that are connected to each other in metabolic and protein interaction networks. Our brain is a network of neurons, and its function emerges through the network, not through the action of individual neurons. But, if you think about it in a wider sense, the social and professional networks that we are part of are a mainstay of our society. We can also talk about economic networks, technological networks, social media, and so on.
What we have understood in the last 20 years is that if we cannot map out the networks behind these systems, we have no chance to understand how they work, because the emerging properties of these systems, their functions are driven not by individual components, but by different components working together in a specific way to achieve certain function.
Network science itself is now a big discipline. It has emerged slightly more than 20 years ago, and now, you can get a PhD in it in Boston, in Vienna, in Budapest. There are thousands of researchers focusing on all kinds of networks. Within biology, however, networks offer a post-genomic path towards understanding disease, because what the human genome has really provided us is the phone book that shows you the components of the cell, but it does not tell you how those pieces interact with each other.
If you ask how the system behaves, the answer is never ‘here’s the list of components’. It’s always, ‘here’s the list of components, and here’s how they interact, which results in certain outcomes and behaviors’. So, if we really want to understand how the cell behaves, and what life is, we have to map the networks, the interactions between the pieces.
Network science aims to further this journey on many levels by creating accurate maps of the subcellular and intracellular environment and developing the mathematical and algorithmic tools that can make sense of these networks. You start from a big network perspective and zoom into a particular disease or a particular gene, you bring the knowledge that is eventually experimentally testable, and this can lead to therapies and cures for disease.
Things like machine learning are probably good instruments to study those networks.
Yes and no. Machine learning is obviously a fabulous tool if you have a giant amount of data, and you need to make sense of it. There are certain aspects of network medicine where machine learning is necessary, but network medicine does not deal with extremely large networks. In the end, we only have 20,000 proteins in our cell. Those, of course, have multiple isoforms, so you’re in a hundred thousand region. You don’t really need machine learning to make sense of that.
What you need machine learning for is to categorize, to classify, and this really is what machine learning does. It finds boxes, puts things into those boxes, and makes predictions based on all this, and yes, it is a useful tool. For example, we use machine learning along with other network medicine tools to predict drug repurposing candidates.
It’s interesting that the 20,000 proteins present in a cell don’t form a giant network. This begs the question, when will we finally have a computer model of a cell?
The question is, what is a model of a cell? There are several models out there. People who’ve been trying to do this got very far away, creating really cool stuff. For instance, they try to take every single interaction and build it into a giant cell model. But when you try to create a model of a cell, it is not enough to have the interactions, you also need to have the dynamics of the interactions.
We have a pretty accurate knowledge of human and bacterial metabolism. Most of the reactions are known. So, we know the network, but we don’t know the kinetic constants. Without having all the kinetic constants, you cannot model the dynamics of metabolism. We have those kinetic constants only for selected pathways, like the cell cycle pathway and a couple of others, but a high-throughput methodology that can determine such kinetic constants in a systematic way is not even on the horizon.
So, do we have the data to map the networks out? Yes. Do we have a way of turning that knowledge into dynamic models? Not yet. Do we want to turn them into dynamical models? Sure. Can we go without that? Yes, and that’s the interesting part. One of the things that we have shown about three years ago, in a paper published in PNAS that was inspired by other people’s work, is that just knowing the network gives you about 70-80% predictive power about the behavior of the system. For the remaining 20-30%, you do need to know the accurate kinetic constants.
Since we don’t even have a hope of getting to know them all, we go as far as we can with what we have, which is the network map, and the idea is that the network maps that the cell has developed within itself are very much attuned by evolution to the function of the system. So, once you know the map, you can deduce most of the functions. Yes, it would be great to know the kinetic constants and make dynamical models, but you can get pretty far in understanding and eventually curing diseases even without that.
At a certain point, you became interested in nutrition. Most of our knowledge in this field comes from population studies, which frankly are not a good source. How do you apply network medicine to nutrition and why is it important?
The Foodome Project, as we nicknamed it in the lab, began about five years ago, when I realized that every network that we had built and curated in the lab is really based on genomic and genetic information, and we know that having just the genetic information gives you only limited predictive power about the occurrence of disease and so on. If you look at heart disease, only about 20% of the occurrence is explained by genetic factors. Where does the rest come from? The environment. So, how could we integrate these environmental effects into the subcellular models that we’ve been building for 15 years?
We started with the food because food is one of the biggest environmental components. There are others, of course, such as stress, exercise, and so on, but the biggest single one is probably food. And we thought that it should be easy because in the end, food is molecules, and molecules get into the cell and interact with various cellular components. So, let’s just figure out what the molecules are and how they interact with the cell, right? That’s when I was in for a big surprise that started this big journey for us. I realized we have absolutely no clue what chemicals are in our food.
What about nutritional science? Sure, it has done a great job of identifying the energy sources and the vitamins that we need to survive. These are typically the chemicals that are processed by our metabolism, like sugars, fats, carbohydrates, and many other things. You must get them on a daily basis because if you don’t, you will not survive.
These are about 150 components, and they are indeed being tracked by the USDA in multiple foods, and we know a lot about them. But to my surprise, already early on, we realized that food has more than 20,000 components. That was four years ago. Today, in my lab, we have a very accurate database of about 135,000 chemicals that can be found in food.
What are these chemicals? We call them the nutritional dark matter because they are not systematically tracked by any agency. We mapped out the full scientific literature, and the knowledge is sporadic. That includes things we hear a lot about, like polyphenols.
Polyphenols can be considered nutritional dark matter because they are not processed by the human metabolism, although there’s some processing by the bacterial metabolism in our gut. Their primary role is not metabolic – they are not a source of energy or of any building blocks in the cell. Their function is rather regulatory. Once they get into the bloodstream and into the cells, they bind to human proteins and regulate their function.
In the end, we have learned that while the nutritional components are essential, they are like the gas and oil for your car, there is also this nutritional dark matter. Amazingly, for two thirds of the molecules that constitute it, we have known health effects. So, the big question is how can we even attempt to understand how our environment affects us, if we don’t have an accurate map and the list of these chemicals?
What the Foodome Project have been doing in the last five years is first, trying to curate an accurate list of all the chemicals that are potentially present in our food and to understand their characteristics. We work with mass spectrometry labs, we comb the literature, we develop tools that predict the presence and the concentration of those chemicals, and that’s one part of the project. The other part is, once you know that the chemical is there, how do you predict its effect on health? Or, if you put it the other way around, we want to pick a disease and find out which food chemicals may have a positive or negative impact on that disease.
This is probably a huge project. Do you have some interesting insights already that you’d like to share with us?
We do have several. First, in our work with polyphenols, we ended up identifying several polyphenols with previously unknown health effects, and connecting one of them to specific health outcomes. We showed that it has an impact on platelet function in the cell, which is relevant for cardiovascular diseases.
The key part is that not only we can determine its impact, we can also precisely predict the pathway, the chemical interactions involved, which makes it experimentally testable. My colleagues at Harvard added this chemical to cell lines and measured the expression patterns of multiple genes and proteins, validating our predictions at a cellular level. So, our methodology allows us to make experimental testable predictions about which chemicals are beneficial or deleterious.
We also predicted what chemicals could affect rheumatoid arthritis, and of the 30 chemicals that we predicted, I believe only one had no effect once it was tested by an independent lab. So, the combination of food and network medicine gives us a pipeline, the ability not only to make experimentally testable mechanistic predictions of how food affects health but also to make disease-specific predictions that may be directly actionable for patients. We are now trying to raise funds for a clinical trial so that we can prove that those chemicals indeed have the impact that we’ve predicted.
This is really interesting considering that nutritionists seem to disagree on some basic things, like the amount of protein that’s good or bad for us. Has your approach added something to this or other discussions that are going on in nutritional science?
Let’s be fair with nutritionists. We consume tens of thousands of chemicals daily, and the challenge that nutritionists have in front of them is to piece out the impact of one or a few chemicals on health. This is such a convoluted problem that it is virtually impossible to solve unless you have millions of patients in clinical trials. So, many of the disagreements in the literature are rooted in the uncertainty or the inability to make accurate measurements of outcomes.
They’re dealing with a massive network-based problem, trying to solve it with standard statistical tools that were developed for Gaussian distributions. There’s a giant gap between the complexity of the problem they’re trying to address and the toolset available to them currently. And I’m amazed at the clean results they were able to get in some cases despite of that, like the impact of sugars and salt.
Or of processed meat?
That’s right. So, if the signal is strong enough, they can discern it. Can we do better? I hope we can. I think an approach based on networks and machine learning, when applied to a large dataset, may give us a way to do this, but we must start designing trials that take advantage of those tools. With the availability of apps where people are tracking their eating pattern and the massive clinical data coming in, I think in the next few years, we’ll see a revolution in the way we understand the impact of food on our health.
I’m looking forward to this revolution. I was also fascinated by your research into processed food. You have developed a measure of just how processed various types of food are, and this measure seems to correlate with health effects. We knew that processed food was bad for you, but this is much deeper. But why actually any amount of processing seems to correlate with health?
We started to understand what processing does to food and how it affects our health only recently. So, let’s start with what is processing. We process food every day, and this isn’t a problem in itself. When you peel an orange, or when you cut a cucumber, salt it, cook it – all this is processing, but those are mechanical and simple chemical processes that do not fundamentally change the chemical composition of the food. Maybe cooking, frying, etc. have some impact, but nothing drastic.
And then there is ultra-processing. It is a precise process that can only be done in factories or laboratories. It’s when you take the food, decompose it, and then reassemble its components into some other product. You are effectively creating new food from food-based components.
Let me give you an example. So, you go to the supermarket, and you buy ‘natural orange juice’. Natural because it’s based on oranges, but it’s ultra-processed because once the oranges are collected, they’re squeezed out. Then, they’re decomposed into three different components, which is the juice itself, the pulp that is taken out, and the water.
Those components are stored separately, and only later, often hundreds of miles away from the collection point, they are reassembled. Whatever you get bears no resemblance to orange juice. It doesn’t taste like it, it doesn’t have the right consistency. So, you must add lots of stuff that give you the consistency, the taste, the smell of orange juice. This is ultra-processed orange juice. What you make at home by squeezing an orange is just processed orange juice.
About five-ten years ago, a Brazilian group has started to systematically classify food in terms of whether it’s ultra-processed or unprocessed. Thanks to their effort, people have started to reanalyze the health data they had on individuals, and to realize that one of the biggest negative health effects is coming from ultra-processed food.
Let me give you some rough numbers. If you move 10% of your calorie intake from unprocessed to ultra-processed food, your chances of diabetes increase by 12%, and your chances of cancer by 10%. There’s a whole list of diseases where the risk suddenly shoots up by 5% to 10%. All you did is replace food that your grandmother would cook with the ultra-processed food that you buy at the supermarket. Otherwise, you’re eating the same things. The reason for that is still a big mystery, but there is already very strong evidence that ultra-processed food has deleterious health effects.
It’s probably not necessarily true about each and every kind of ultra-processed food, it’s just an average, right?
We don’t know that because the clinical trials so far just say what percentage of your calories is coming from ultra-processed food and whether this correlates with health outcomes, but people can go deeper. It has been shown that when it comes to meat, most of the health effects that we previously assigned to red meat are limited to ultra-processed meat. What’s important is that in the US, calorie intake is dominated by processed food. Americans get 60% of their calories from ultra-processed food. That’s mind-blowing, and that’s where the problem is.
I wonder if this has something to do with the obesity epidemic.
Yes, of course. This is the other result that clearly comes out of the data: that moving 10% of your calories from less-processed to ultra-processed food increases your chances of developing obesity and metabolic syndrome by the same 10%. So, most of the bad health outcomes that are systemic in the American population can be linked to consumption of ultra-processed food.
Here’s one problem with ultra-processed food that we come in to solve. When you go to the supermarket, you have no idea whether the food that you’re about to buy is ultra-processed or merely processed. That information is not available anywhere on the boxing or in databases. We built an AI tool that looks at all the foods at the supermarket, and we have created a website called Truefood.tech where you can type in your favorite food, and it will tell you the degree of processing it was subjected to. It will also tell you what other items of the same type are sold in the same store, that are less processed, so it offers you an alternative.
That came out of our foodome research. We were curious about not only what chemicals are in the food but also what is the concentration of the chemicals that are in the food? We made a very surprising discovery that was just recently published in Nature Food. We realized that in natural foods, the variation in the concentration of specific chemicals is very narrowly bounded, and there’s a precise mathematical formula that describes, say, how much vitamin C and how much sugar is expected to be in your food.
When we apply the same formula to ultra-processed food, we see major deviations because when we reassemble food from components, we don’t respect the natural concentrations anymore. Rather, companies do whatever they think is right taste-wise or consistency-wise. So, we create a chimera that has the same ingredients as the original food and then some, but in different proportions.
Why is that a problem? Let’s say I give you 10 atoms of hydrogen and one atom of oxygen and ask you to make me a lot of water. You can’t because one atom of oxygen can only take up two atoms of hydrogen, and the rest is just floating around. Previously we thought that getting all those extra chemicals with food was no problem because they would just go in and out, the body doesn’t know what to do with them. But the body has been evolutionary engineered to deposit anything it gets because you never know when it’s going to become useful. In the end, much of this stuff gets deposited, like all the cholesterol in your arteries.
This stuff comes into your body through many different foods, and the body doesn’t know what to get rid of. Our hypothesis was that the problem with ultra-processed food it that it is chemically unbalanced. We normally eat ourselves, meaning that anything we eat has the same carbon-based chemical engine as the human metabolism. But ultra-processed food is different, the ratios of chemicals are different.
So, our AI model looks at the nutritional components that every food producer must disclose by law, and it detects these fine changes in the concentrations and determines how ultra-processed the food is. You can’t tell it by just looking at the label. Yes, maybe it has a little bit more sugar, maybe a little more salt or carbohydrates, but you don’t know what the proper ratio should be. The AI, on the other hand, learns and knows, and immediately detects if the food has been tampered with.
I think this online tool that you built is extremely useful, although I was mostly reassured by it that I’m doing okay. One food that is hard to replace is bread, and all bread is processed to some extent.
Bread does not need to be heavily processed. It’s a simple combination of yeast, grain, and a couple of other things. Why do we ultra-process bread? Because making bread at home, that is, by processing, and not ultra-processing, takes a day, but if you want to make large amounts of it fast, you must add new processes.
Even at the supermarket, as our database shows, you can find minimally processed bread beside ultra-processed bread, and the same goes for most foods. Yes, for some food categories, ultra-processing is necessary, but for most, there is a minimally processed alternative.
Why is this important? As we show in our paper, if you just take your normal eating pattern (we eat about 20 different items per day), and you replace one item with a less-processed version, this can already give you a considerable reduction in many of the outcomes associated with ultra-processed food, because typically, there’s always one item in your diet that is the biggest contributor to your ultra-processed food intake.
Just replace it. Keep eating the same. If it’s a burger, replace it with an unprocessed burger. If it’s soda, replace it with water, and so on. This single change that just replaces one item with another without changing your diet could have a huge impact on your health.
Yes, this particular parameter blew me away when I read your paper. You also mention the obvious fact that processed food is cheaper. So, basically, eating less processed food can be considered a privilege that may not be available to everyone.
Yes, of course. Ultra-processed food has clear advantages: it’s cheaper, it has a longer shelf time, which is also reflected in the price, it has higher caloric content, and so on. Do you know where processed food comes from? It turns out that the story goes back to a place about 50 miles from my home here in Boston. It was originally developed by the US military that needed non-perishing food with high energy content, light enough for the soldiers to carry it on the battlefield. In the 1960-70s, Congress allowed food companies to start utilizing the patents developed by the military, and that’s when ultra-processed food invaded American supermarkets.
I think that if we care about population health, we will find a way to get less processed food to people, but we also need to know what is processed and what is not, and there has to be awareness and demand. There’s a lot you can do with tax policy to encourage consumption of less processed food over ultra-processed food, and many European countries are already doing that. I think eventually, the solution will come not from simply banning processed food but from doing what we did with sugary food and cigarettes, which is to tax the hell out of it and use the money to promote and support the production of healthier food. It is possible, but it’s a long-term journey, and it requires legislation and leadership.
Pandemics are also networks, right? What can network science teach us about the COVID pandemic and pandemics as a whole?
Sure, epidemics are network-based problems. Viruses spread through social networks, and that’s why the current leader of my Network Science Institute in Boston, Alessandro Vespignani, has also been the White House modeler of COVID since March 2020. He has predicted the problems that we’re going to have with COVID already in December 2019, way before the pandemic really took off. Many other network scientists are involved in predicting COVID patterns and continue to study it. I would go as far as to say that there’s no way you can build good predictive tools for disease spreading without considering the network perspective.
Curing diseases is also fundamentally a network problem. My lab has been deeply involved in the fight against COVID from early on, in order to identify, using tools from network medicine, drugs that could be repurposed for COVID patients, because at that time, it wasn’t clear how long creating vaccines will take, and the only solution was to repurpose existing drugs. We ended up predicting and testing, together with colleagues at Boston university, 6,000 drugs and identifying about a dozen that could be, and were, repurposed for COVID patients.
Fundamentally, for any infectious disease, the first person you need to consult is a network epidemiologist. That’s what’s happening right now with the outbreak of monkeypox. You probably can google it and find out that many of my colleagues are already involved in modeling the spread of this disease. They were also involved in the modeling of Ebola and Zika, and so on. There’s no way to understand and stop infectious diseases without a network perspective.
How would you describe the general situation in the longevity field today? What are you excited or frustrated about, and what directions do you think are especially worth pursuing?
I think that longevity research is undergoing a fundamental transformation right now. The pieces have been assembled, and the will is there to really try to make an impact. However, one of the challenges with longevity is that everybody’s hoping for that simple pill, but also everybody who is really in the field understands that the aging is not a single mechanism; there are multiple components and mechanisms involved. Aging is the sum of many different diseases and conditions that come together, and they are always coupled with each other.
So, I believe, and that’s what we are focusing on together with Vadim Gladyshev, that we must bring the network toolbox to aging research. I think this is the only way to put those many pieces together and perhaps find some solutions, although we are far from actually seeking for solutions right now; we’re trying to map out the networks involved in longevity and aging.
If we are successful at that, then we can also think of potential interventions that may alter those pathways in the way we’re doing with other diseases. The bottom line is, based on what I see by reading the literature and going to conferences, every field has this moment when it totally rejuvenates itself, if we use our favorite term, and I feel like this moment for longevity research has arrived.